Python Data Skills 15: Automated Data Cleaning Part 2
Imputation, Automated GroupBy + Mean Imputation, Automated Timeseries Imputation, Automated KNN Imputation
In the previous post we talked about how we can automatically spot our outliers. More specifically, automatically knowing which column & which row number they are in.
Spotting outliers is great because you can use them for visualization purposes, reports, anomaly detection, etc…
But, in this post, let’s say you've taken interest in cleaning them out. Usually, it is far easier to just straight up remove the outlier, replace it with a zero, replace it with a mean, etc... But, in this case, I’m going to show you can you can replace that outlier with it’s most sensible value, using imputation.
Note: If you have the functions from part 1, you can use those. This is by getting them to replace the outlier value with an NA. Afterwards, you can use the functions presented here. These functions can automatically perform imputation on them.
Table of Contents
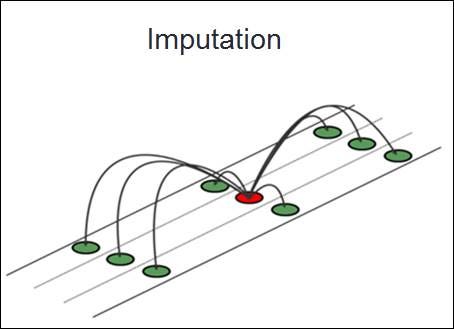
Imputation
Automated GroupBy + Mean Imputation
Automated Timeseries Imputation
Automated KNN Imputation
1 - Imputation
Handling missing values in a dataset is a fundamental step in the data cleaning process. Leaving missing values unattended can mislead analyses and distort final results. A very common strategy for dealing with these missing values is imputation. Imputation is the process of substituting missing data with estimated values.
1.1 What is Imputation
Imputation is the method of filling in missing data points in a dataset with some estimated values. The goal is to preserve the dataset's overall integrity, ensuring that statistical analyses and model training remain unbiased. Imputation is a vital tool in a data scientist's arsenal, as real-world data often contains gaps. Instead of eliminating rows with missing data – which can result in a loss of valuable information – we fill in these gaps, or "impute" them.
1.2 Mean/Median Imputation
One imputation method replaces missing values with the column's mean or median. This is one of the simplest and most common imputation methods. This method is particularly useful for features with numerical data:
Mean Imputation: Replace missing values with the average of the available data. This is best suited for data that is normally distributed.
Median Imputation: Use the median (the middle value in an ordered list) for imputation. This is typically used for data that has outliers or is not normally distributed. This is because in these instances, the median is less sensitive to extreme values than the mean.
import pandas as pd
def impute_missing_values(df, column, strategy="mean"):
if strategy == "mean":
df[column].fillna(df[column].mean(), inplace=True)
elif strategy == "median":
df[column].fillna(df[column].median(), inplace=True)
else:
raise ValueError("Strategy should be 'mean' or 'median'.")
return df
# Usage:
imputed_df = impute_missing_values(my_dataframe, "column_name", strategy="median")The fillna method in pandas provides a streamlined way to replace NaN (NULL) values. This is within a specified column with either its mean or median.
2 - Automated GroupBy + Mean Imputation
Imputation plays a pivotal role in managing missing data. Methods such as replacing NAs with mean or median are quick. But they often oversimplify the underlying patterns of the dataset. Leveraging the relationships between columns is beneficial. It can elevate the integrity and reliability of the data. Adding a touch of granularity to imputation processes can also do this.
2.1 Limitations of Mean/Median Imputation
The primary limitation of replacing missing values with the column's mean or median is its sweeping generalization. It assumes all missing values, regardless of other features or conditions in the dataset, share the same inherent characteristics, which is rarely the case. This can distort the distribution of values, potentially introducing biases in analyses and predictive modeling.
2.2 Exploring Correlation for insight
Before imputation, understanding relationships between columns is key. Correlation analysis provides insights into how one variable moves in relation to another. If two columns, say col1 and col2, are strongly correlated. These two columns can share a systematic relationship. This means we want to harness this during imputation.
In this example, I’ll assume you are trying to perform imputation on col1:
correlation_matrix = df.corr()
print(correlation_matrix['col1'])2.3 Binning for granularity
To improve the accuracy of imputation, consider segmenting correlated columns into bins or groups. This process involves categorizing a continuous variable based on ranges or intervals. For instance, if col1 has missing values and is correlated with col2, segmenting the values of col2 can aid in better imputation for col1.
df['col2_bins'] = pd.cut(df['col2'], bins=5, labels=['very low', 'low', 'medium', 'high', 'very high'])2.4 GroupBy + Mean Imputation
With the data segmented, it's possible to replace missing values in col1 based on the mean of their respective bins from col2. This ensures imputation is more sensitive to the nuances in data.
Now… let’s be real doing all of the above steps is hard, so, wouldn’t it be great if you had a function, and it would basically do the entire thing for ya?
Anyways, here’s a function that I store in my personal github that does the entire thing for ya:
Keep reading with a 7-day free trial
Subscribe to Data Science & Machine Learning 101 to keep reading this post and get 7 days of free access to the full post archives.