Python Data Skills 16: Miscellaneous Cleanup
Dimensionality Reduction, Encoding Your Data, Sparse Matrix, Data Leakage
We’ve spent a huge amount of time wrangling data, and cleaning the absolute fk out if it. This is what you do in the real world btw. But there were a few minor topics I had missed. And, I intend to cover them here, these are simple minor definition, and topics. You can think of this as kind of like a glossary at the end.
Table of Contents
Dimensionality Reduction
Encoding Your Data
Sparse Matrix
Data Leakage
1 - Dimensionality Reduction
Dimensionality reduction is a technique employed to reduce the number of features or variables in a dataset while trying to preserve as much information as possible. It can be categorized as a key step in the data preprocessing pipeline, especially when working with datasets that have a large number of features or columns.
1.1 Why do we do Dimensionality Reduction
Here’s a list of reasons why we reduce the dimensions (number of columns) in our data:
Computational Efficiency: Reducing dimensions can lead to a significant decrease in storage and computation times.
Visualization: Visualizing high-dimensional data can be challenging. Dimensionality reduction techniques allow us to reduce the number of dimensions, making it possible to visualize the data in 2D or 3D space.
Avoid Overfitting: With a high number of features and insufficient training data, a model may overfit. By reducing dimensions, we can mitigate the risk of overfitting.
Improved Performance: In some cases, removing noise and redundancy can lead to improved model performance.
1.2 Principal Component Analysis (PCA)
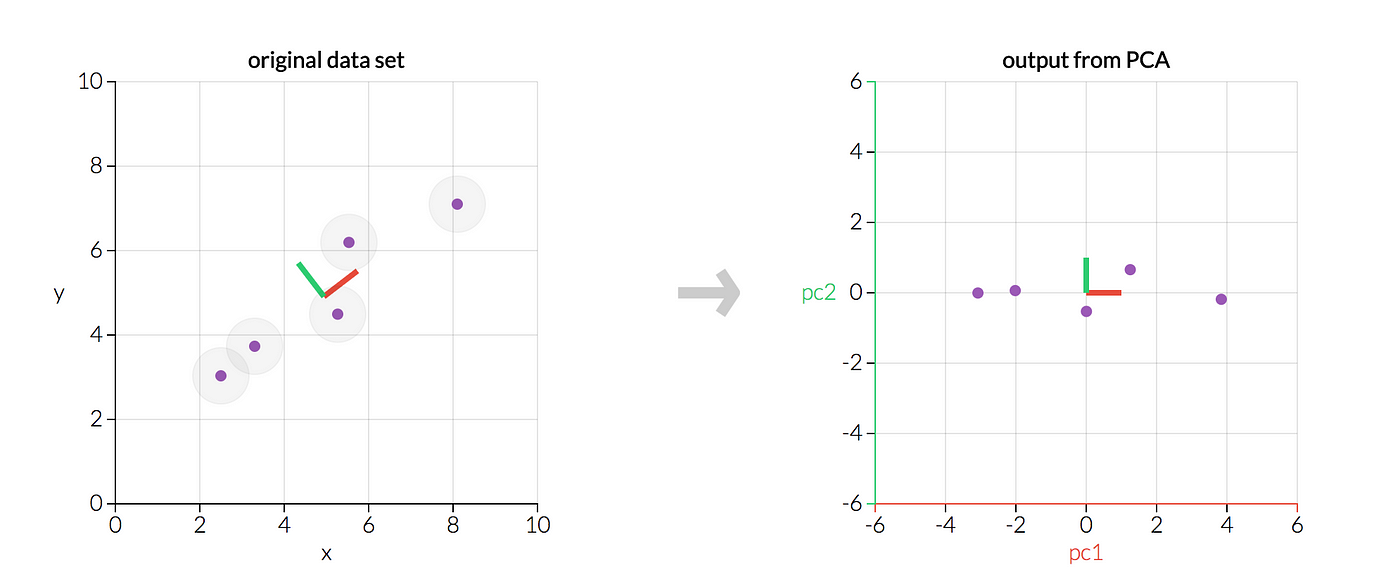
PCA is a popular technique for dimensionality reduction. It identifies the axes (or "principal components") in the data that maximize variance. These axes are orthogonal to each other. This means they're perpendicular in the n-dimensional space. Once identified, we can project our original data onto these new axes. This will reduce the number of dimensions.
Steps to perform PCA:
Standardize the dataset (mean of 0 and variance of 1 for each feature).
Compute the covariance matrix of the standardized data.
Obtain the eigenvalues and eigenvectors of the covariance matrix.
Sort eigenvectors by decreasing eigenvalues and choose the first 'k' eigenvectors (where 'k' is the number of dimensions you want your data to be reduced to).
Transform the original dataset through its projection to the selected 'k' eigenvectors.
from sklearn.decomposition import PCA
from sklearn.preprocessing import StandardScaler
def perform_pca(data, n_components=5):
# Step 1: Standardize the dataset
scaler = StandardScaler()
data_std = scaler.fit_transform(data)
# Step 2 to 5: Apply PCA
pca = PCA(n_components=n_components)
transformed_data = pca.fit_transform(data_std)
return transformed_data
# Example usage:
reduced_data = perform_pca(your_dataframe, n_components=3)
Note: If you are working with time series data, you have to be careful. You have to train the PCA on your training data. Then perform it on your test data or you might encounter data leakage (more info on it below)
1.3 Why PCA is gold standard for Dimensionality Reduction
Versatility: PCA can be applied to almost any dataset and doesn't make strong assumptions about the structure of the data.
Orthogonal Transformation: The principal components produced by PCA are all orthogonal to each other, ensuring no redundant information.
Maximizing Variance: PCA focuses on maximizing variance, ensuring that the most informative features are retained.
Ease of Interpretation: Once dimensions are reduced using PCA, the resulting principal components often allow for easier interpretation and visualization.
One limitation of PCA is that it's a linear technique. It may not capture complex nonlinear relationships in the data.
2 - Encoding Your Data
2.1 Understanding Categorical Data
Categorical data, as the name implies, represents categories. These are data points that can take on one of a limited set of values. They are especially common in datasets where we have labels. They are also common in data sets with identifiers or non-numeric features. Categorical data are generally classified into two types:
Nominal: These are categories that don't have a natural order. Examples include colors (red, blue, green) or cities (London, Paris, Tokyo).
Ordinal: These categories have a clear, distinct ordering. For instance, ratings such as low, medium, high, or educational levels like primary, secondary, and tertiary are ordinal in nature.
Encoding is essential because many machine learning algorithms need numeric input features. They cannot handle categorical data directly. This can be in linear regression or support vector machines.
2.2 Dummy Variable Encoding
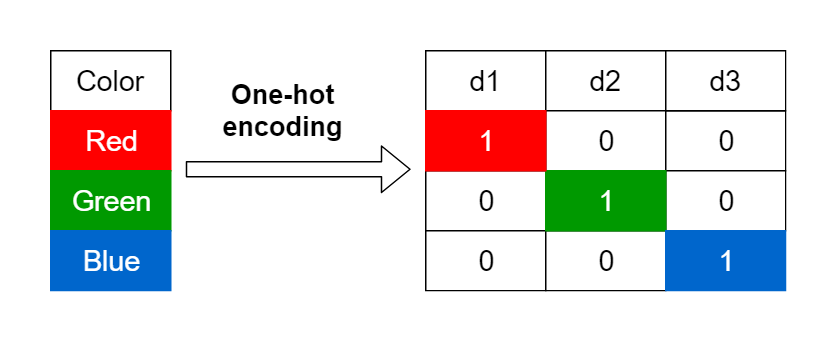
For nominal data, one of the most common forms of encoding is creating dummy variables. This is often known as "one-hot encoding." This process transforms a single column of categorical data into multiple columns. One for each category. For instance, a single column with colors red, blue, and green becomes three columns:
A column that has a 1 if the color is red and 0 otherwise.
A column that has a 1 if the color is blue and 0 otherwise.
A column that has a 1 if the color is green and 0 otherwise.
The main advantage of this method is that it doesn't introduce any arbitrary numbers that the model could misinterpret as ordinal.
2.3 Ordinal Data Encoding
For ordinal data, where categories have an inherent order, it makes sense to convert categories into numbers that preserve this order. For example, ratings of low, medium, high could be encoded as 1, 2, 3, respectively.
However, care must be taken to ensure that the encoding captures the true relationship between categories. Assigning arbitrary numbers could lead to incorrect models and interpretations.
2.4 Python Function for ya
You can use this function to take care of the encoding for ya:
import pandas as pd
def encode_data(data, column_name, encoding_type='dummy', order_list=None):
"""
Encodes a column in a dataframe.
Parameters:
- data: DataFrame, input data
- column_name: str, name of the column to be encoded
- encoding_type: str, either 'dummy' or 'ordinal'
- order_list: list (optional), order of categories for ordinal encoding
Returns:
- DataFrame with the encoded data
"""
if encoding_type == 'dummy':
# Get dummy variables and drop the original column
dummies = pd.get_dummies(data[column_name], prefix=column_name)
data = pd.concat([data, dummies], axis=1)
data.drop(column_name, axis=1, inplace=True)
elif encoding_type == 'ordinal' and order_list:
cat_dtype = pd.CategoricalDtype(categories=order_list, ordered=True)
data[column_name] = data[column_name].astype(cat_dtype).cat.codes
return data
# Example usage:
# For dummy encoding
# encoded_data = encode_data(your_dataframe, 'color_column')
# For ordinal encoding
# encoded_data = encode_data(your_dataframe, 'rating_column', encoding_type='ordinal', order_list=['low', 'medium', 'high'])3 - Sparse Matrix
3.1 Understanding a Sparse Matrix
A matrix is considered sparse if a large proportion of its elements are zeros (or, more generally, the default fill-value). In contrast, a matrix is said to be dense if most of its values are non-zero. Sparse matrices arise naturally in many real-world problems, particularly in situations where we're dealing with large datasets with missing or unobserved values, or data that inherently contains lots of gaps.
Imagine a user-movie rating matrix in a movie recommendation system. Not all users rate every movie, resulting in a matrix full of unobserved values, making it sparse.
3.2 Why use a sparse matrix
Memory Efficiency: Storing all those zeros in a dense matrix is wasteful. A sparse matrix only stores non-zero elements, saving significant memory. For very large matrices, converting to sparse format can be the difference between being able to fit the matrix into memory or not.
Computational Efficiency: Algorithms can be adapted to exploit the sparse structure and become faster as they don't need to process zero values.
Data Compression: Sparse formats can significantly reduce the storage requirements for datasets.
However, there's a trade-off. While operations like matrix-vector multiplication can be faster with sparse matrices, some operations can be slower due to the overhead of accessing the sparse structure.
3.3 Converting a DF to a sparse matrix
Pandas provides functionality to deal with sparse data structures. Convert it to a sparse structure, reducing memory usage. You can do this when you have a DataFrame with lots of missing or zero values.
Let's look at how we can do this and a custom function to aid the process:
import pandas as pd
from scipy.sparse import csr_matrix
def convert_to_sparse(df):
return csr_matrix(df.fillna(0).values)
# Example usage:
matrix_sparse = convert_to_sparse(your_dataframe)4 - Data Leakage
Data leakage, in the context of machine learning and data science, refers to a mistake made during the preprocessing or training phase where information from the testing dataset is used inappropriately before or during training. In simpler terms, it's when the model accidentally gets a hint about the testing data. This gives the illusion that a model performs exceptionally well, but in reality, when applied to unseen data, the performance can be abysmal. This can happen in various ways, such as including the test data in the training dataset, or using features that indirectly have information about the target variable.
4.1 Why is data leakage important?
Misleading Performance Metrics: Data leakage can lead to overly optimistic performance metrics. This might make one think that the developed model is ready for production, but its real-world performance might be unsatisfactory.
Wasted Resources: Training models based on leaked data might result in wasting computational resources and time.
Poor Generalization: Since the model never truly learned the underlying patterns of the data (it simply "cheated" with the leaked data), it would likely perform poorly on genuinely unseen data.
Business Consequences: In industries like finance or healthcare, poor model performance can lead to significant financial losses or critical errors.
4.2 Solutions to data leakage
Separation of Data: Ensure that the testing dataset is kept separate from the training dataset at all times. This might sound elementary, but accidental mix-ups can occur.
Temporal Split: In time-series data, always use past data to predict future events to ensure that no future information leaks into the past.
Care with Feature Engineering: Be wary of features that might indirectly give away information about the target variable. For instance, if you're trying to predict a health outcome, including a feature like 'amount of medication taken' might indirectly give away the severity of the patient's condition.
Use Pipelines: In Python's
scikit-learn, using pipelines can prevent data leakage during the data preprocessing phase. It ensures that operations like scaling or normalization are done separately on training and testing data.Cross-Validation: Implement proper cross-validation schemes to ensure that the validation phase is robust and leakage-free.
Review Data Sources: Ensure that there's no overlap between datasets, especially when merging data from different sources.
This piece of code can help you avoid data leakage:
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
def build_pipeline(data, target_column):
X = data.drop(target_column, axis=1)
y = data[target_column]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
pipeline = Pipeline([
('scaler', StandardScaler()),
('classifier', RandomForestClassifier())
])
pipeline.fit(X_train, y_train)
return pipeline
# Usage example:
trained_pipeline = build_pipeline(your_dataframe, "your_target_column")In the function above, the pipeline ensures that StandardScaler scales the data separately for training and testing, thereby avoiding leakage.